GMDH-type neural network algorithm is a heuristic self-organizing algorithm to model complex systems. This ultimate guide involves feature selection and classification via GMDH algorithm for a binary response. Find out how to apply GMDH algorithm in R.
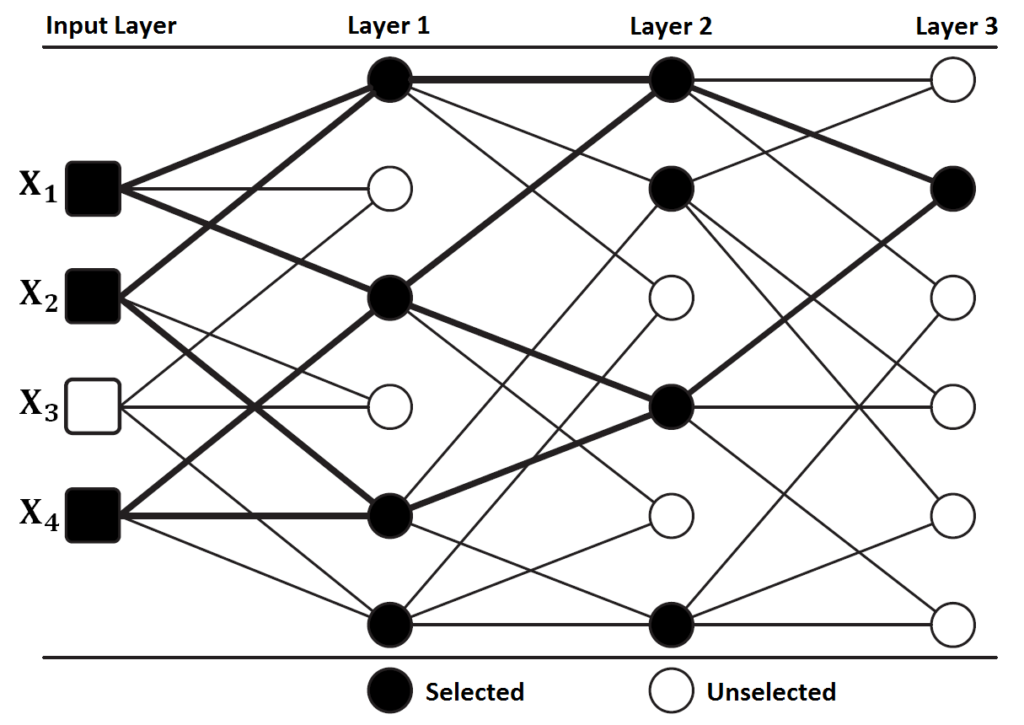
In this tutorial, we will work GMDH-type neural network approach for feature selection and classification when a response with two classes exists. Before we start, we need to divide data into three parts; train, validation and test sets. We use train set for model building. We utilize validation set for neuron selection. Last, we show the performance of the model on test set.
Check Out: How to Recode Character Variables in R
In this tutorial, we will implement the algorithm on breast cancer dataset, also used in the work done by Dag et al. (2019), available in mlbench package (Leisch and Dimitriadou, 2010). Before we go ahead, we load dataset and start to process the data.
library(mlbench)
data(BreastCancer)
After loading dataset, let’s exclude missing values to work on the complete dataset.
data <- na.exclude(BreastCancer)
head(data)
## Id Cl.thickness Cell.size Cell.shape Marg.adhesion Epith.c.size Bare.nuclei Bl.cromatin Normal.nucleoli Mitoses Class
## 1 1000025 5 1 1 1 2 1 3 1 1 benign
## 2 1002945 5 4 4 5 7 10 3 2 1 benign
## 3 1015425 3 1 1 1 2 2 3 1 1 benign
## 4 1016277 6 8 8 1 3 4 3 7 1 benign
## 5 1017023 4 1 1 3 2 1 3 1 1 benign
## 6 1017122 8 10 10 8 7 10 9 7 1 malignant
Also Check: How to Handle Missing Values in R
We need to define the output variable as vector and input variables as matrix.
x <- data.matrix(data[,2:10])
y <- data[,11]
We need to divide data into three sets; train (60%), validation (20%) and test (20%) sets. For reproducibility of results, let’s fix the seed number to 100. Then, we obtain the number of observations in each fold.
set.seed(100)
nobs <- length(y)
ntrain <- round(nobs*0.6,0)
nvalid <- round(nobs*0.2,0)
ntest <- nobs-(ntrain+nvalid)
Now let’s obtain the indices of train, validatation and test sets. Before we obtain the indices, we shuffle the indices to prevent any bias based on order.
indices <- sample(1:nobs)
train.indices <- sort(indices[1:ntrain])
valid.indices <- sort(indices[(ntrain+1):(ntrain+nvalid)])
test.indices <- sort(indices[(ntrain+nvalid+1):nobs])
We can construct train, validatation and test sets.
x.train <- x[train.indices,]
y.train <- y[train.indices]
x.valid <- x[valid.indices,]
y.valid <- y[valid.indices]
x.test <- x[test.indices,]
y.test <- y[test.indices]
After obtaining train, validation and test sets, we can use GMDH-type neural network algorithm. GMDH algorithm is available in GMDH2 package (Dag et al., 2019).
library(GMDH2)
model <- GMDH(x.train, y.train, x.valid, y.valid)
##
## Structure :
##
## Layer Neurons Selected neurons Min MSE
## 1 36 15 0.0362031658826178
## 2 105 15 0.0311522877861282
## 3 105 15 0.0292792665461301
## 4 105 15 0.0290895694619813
## 5 105 15 0.0290462745341987
## 6 105 15 0.0290366012593365
## 7 105 15 0.0290328987038381
## 8 105 1 0.0290327200978176
##
## External criterion : Mean Square Error
##
## Feature selection : 6 out of 9 variables are selected.
##
## Cl.thickness
## Cell.size
## Cell.shape
## Marg.adhesion
## Epith.c.size
## Bare.nuclei
Also Check: How to Clean Data in R
Now, let’s obtain performance measures on test set.
y.test_pred <- predict(model, x.test)
confMat(y.test_pred, y.test, positive = "malignant")
##
## Confusion Matrix and Statistics
##
## reference
## data malignant benign
## malignant 41 2
## benign 5 88
##
##
## Accuracy : 0.9485
## No Information Rate : 0.6618
## Kappa : 0.8832
## Matthews Corr Coef : 0.8843
## Sensitivity : 0.8913
## Specificity : 0.9778
## Positive Pred Value : 0.9535
## Negative Pred Value : 0.9462
## Prevalence : 0.3382
## Balanced Accuracy : 0.9345
## Youden Index : 0.8691
## Detection Rate : 0.3015
## Detection Prevalence : 0.3162
## Precision : 0.9535
## Recall : 0.8913
## F1 : 0.9213
##
## Positive Class : malignant
Accuracy of GMDH algorithm is estimated to be 0.9485. This algorithm classifies 94.85% of persons in a correct class. Also, sensitivity and specificity are calculated as 0.8913 and 0.9778. The algorithm classifies 89.13% of the persons having breast cancer, 97.78% of the persons not having breast cancer.
The application of the codes is available in our youtube channel below.
Don’t forget to check: 6 Ways of Subsetting Data in R
References
Dag, O., Karabulut, E., Alpar, R. (2019). GMDH2: Binary Classification via GMDH-Type Neural Network Algorithms – R Package and Web-Based Tool. International Journal of Computational Intelligence Systems, 12:2, 649-660.
Leisch, F., Dimitriadou, E. (2010). mlbench: Machine Learning Benchmark Problems. R package version 2.1-1.

0 Comments
1 Pingback